Content
Signal Peptide Peptidase (SPP) and their homologues (SPPLs)
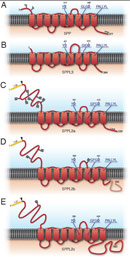
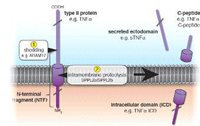
The concept of proteases cleaving their substrates within the hydrophobic core of their transmembrane domains is very well established. In the past fifteen years, various intramembrane cleaving proteases (I-CLiPs) and their respective substrates have been identified. Signal peptide peptidase (SPP) and its homologues, the signal peptide peptidase-like proteases (SPPLs) are aspartyl I-CLiPs of the GxGD-type. In humans the SPP/SPPL family consists of five members: SPP, SPPL2a, SPPL2b SPPL2c and SPPL3 (Fig. 1). SPP and SPPLs are closely related to presenilins, which form the active subunit of the γ-secretase complex. All members of this protease family share a conserved GxGD motif within their catalytic centre. GxGD proteases have turned out to be part of a fundamental sequential proteolytic processing pathway of single span transmembrane proteins, termed regulated intramembrane proteolysis (RIP). Substrates of RIP are typically cleaved within their luminal domain to release a large part of their ectodomain or to clip a hairpin loop between two transmembrane domains (TMD). This cleavage is termed “shedding” and generates a membrane-retained fragment, which is subsequently endoproteolysed within the lipid bilayer by an I-CLiP, causing the release of protein fragments into the cytosol (intracellular domains, ICDs) and the lumen/ extracellular space (C-peptides) (Fig. 2).
While a genetic link between the γ-secretase complex and familial cases of Alzheimer‘s disease is well-documented, the SPP/SPPL family has only recently been identified and their likely diverse physiological functions as well as their regulation have not yet been completely unravelled.
Our research, therefore, aims to unravel the physiological function of the SPP/SPPL-family members and to understand how SPP/SPPL proteases mechanistically select and cleave their substrates within the membrane.